
Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
1
How to cite this article:
Majikes JM, Liddle JA (2021) DNA Origami Design: A How-To Tutorial.
J Res Natl Inst Stan 126:126001. https://doi.org/10.6028/jres.126.001
DNA Origami Design: A How-To Tutorial
Jacob M. Majikes and J. Alexander Liddle
National Institute of Standards and Technology,
Gaithersburg, MD 20899, USA
Jacob.Majikes@nist.gov
Liddle@nist.gov
While the design and assembly of DNA origami are straightforward, its relative novelty as a nanofabrication technique means that the
tools and methods for designing new structures have not been codified as well as they have for more mature technologies, such as
integrated circuits. While design approaches cannot be truly formalized until design-property relationships are fully understood, this
document attempts to provide a step-by-step guide to designing DNA origami nanostructures using the tools available at the current
state of the art.
Key words: DNA nanofabrication; DNA origami; self-assembly.
Accepted: November 27, 2020
Published: January 8, 2021
https://doi.org/10.6028/jres.126.001
1. Introduction
Deoxyribonucleic acid (DNA) origami is a powerful approach for fabricating nanostructures with
molecular precision over length scales of approximately 100 nm. In addition, the ability to functionalize
nanoscale objects with DNA, and other attachment chemistries, means that origami can be used as a
molecular “breadboard” to organize heterogeneous collections of items such as biomolecules, carbon
nanotubes, quantum dots, gold nanoparticles, fluorophores, etc.
On completing this tutorial, readers will have learned how to apply a basic design approach for developing
DNA origami nanostructures. Since the design-property relationships of such nanostructures are still a subject of
active research, we have chosen to focus on basic concepts, rather than trying to create a comprehensive guide.
1.1 Audience
This tutorial is designed to be used by novice designers of DNA origami systems, particularly those
performing undergraduate or early graduate-level research.
1.2 Education or Skill Level
Readers of this tutorial should be familiar with the physical properties of B-DNA, single-stranded
DNA (ssDNA), and crossover junctions. In addition, once ready to create a structure for a specific
application, the designer should determine the full list of functional requirements. This list includes
answers to the following questions: What should the structure do? What specific properties are critical to
the system’s performance?

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
2 https://doi.org/10.6028/jres.126.001
1.3 Prerequisites
The designer should have either sufficient paper for manual design (not recommended) or a design
program such as cadnano [1] (all versions sufficient), nanoengineer
®
, Parabon inSēquio
®
, or equivalent.
1
A
registered account with three-dimensional (3D) structure prediction servers such as CanDo [2, 3] is also
recommended.
1.4 Tools or Equipment
Equipment includes desktop or laptop computer equipment, craft supplies for macroscale models, and
DNA nanotechnology computer-aided design (CAD) software.
1.5 Background
The sequence specificity associated with Watson-Crick-Franklin base pairing in DNA and the ability
to synthesize DNA in an arbitrary sequence [4] combine with the ease of labeling other functional
nanomaterials with DNA to make DNA an ideal molecular Lego
®
. It can be used to create complex
nanostructures in large quantities at low cost, and it has capabilities that are complementary to top-down
nanostructure fabrication methods. However, the design approach required to design and deploy this
molecular assembly tool is different compared to those used for established top-down assembly techniques.
The structure of B-form double-stranded DNA (dsDNA) is shown in Fig. 1. While the structure of
DNA is a surprisingly rich field of study [5], only a few properties are relevant for programmed self-
assembly [6]. These relate to the details of the lock-and-key base-pair interaction between the adenine and
thymine (A-T) and guanine and cytosine (G-C) bases in DNA polymers. Due to the asymmetry of the DNA
polymer and the shapes of the bases, canonical base pairs only form between antiparallel strands of DNA.
DNA strands can be described as antiparallel because each strand has a chemical direction, represented by
the asymmetric carbon positions (5′ and 3′) at which the phosphates are bonded to the sugar. By
convention, the start of the DNA strand is called the 5′ end, because it terminates with a phosphate at the 5′
carbon position, while the final base of the strand is called the 3′ end, because it terminates with a hydroxyl
group at the 3′ carbon position. Also, because the base-pair geometry is not symmetric, the helix has a
minor groove and a broader major groove. If a strand were to leave the helix, e.g., to make an addressable
sticky end, it would be connected to the helix at the last phosphate/sugar moiety attached to a base engaged
in the helix. As such, whether in crossovers or sticky ends, the choice of the z-axis position for a connection
to the helix determines the rotation of that position around the helix.
The principal mechanism of nanostructure fabrication using DNA relies on the ability of DNA strands to
cross over from one dsDNA helix to another adjacent helix, thereby connecting them. These “crossovers” can
be programmed to occur at specific locations using sequence design and base pairing of ssDNA strands to
create dsDNA helices. Crossovers may be formed anywhere, subject to the constraint that the phosphate
backbones are aligned. Figure 2 shows the detailed helix and schematic line representations of a DNA
crossover. One-dimensional (1D) self-assembly along the helix can be used to create both two-dimensional
(2D) and 3D structures. The reader is directed to numerous recent reviews for structure examples [7, 8].
Side-by-side comparison of crossover representations is useful because the design process occurs using
the line representation, but the rules allowing crossovers arise from physical realities visible in the helical
representation. Additionally, one should note that the crossovers shown in Fig. 2 are antiparallel. That is to
say, the yellow and green strands reverse their direction on being forced to exchange helices. While
1
Certain commercial items are identified in this paper in order to specify the experimental procedure adequately. Such identification
does not imply recommendation or endorsement by NIST, nor does it imply that the software, materials, or equipment identified are
necessarily the best available for the purpose.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
3 https://doi.org/10.6028/jres.126.001
structures have been designed with parallel crossovers where the strands do not reverse [6, 9], they have not
readily been allowed by most CAD tools and represent a minority of crossovers used.
Fig 1. Structure of B-form dsDNA and major/minor grooves arising from base-pair geometry in which the sugar/phosphates split the
head-on view of the double helix into 3.6 rad (210 °) and 2.6 rad (150 °), respectively. The phosphate groups that link the sugars of
bases in the z axis are in and out of page in this illustration. The major and minor grooves here are not to scale.
Fig. 2. DNA crossover between helices. (Left) Helix representation. (Middle) Base pairing. (Right) Schematic line representation
showing antiparallel base pairing with 5′ and 3′ ends denoted by squares and triangles, respectively.
While a full review of DNA self-assembly techniques is beyond the scope of this document, we
illustrate how typical structures can be built from connected crossovers in Fig. 3. DNA tiles (Fig. 3A) often
require the fewest distinct DNA strands, and can polymerize in a variety of ways [6]. They are, however,
notoriously sensitive to design and stoichiometry, and implementation of a tile system often requires time-
consuming tuning of relative strand concentrations [6]. Developed, in part, to overcome this problem, DNA
origami (Fig. 3B) comprise a long scaffold, derived from a viral genome, held together by crossovers
programmed into synthetic ssDNA strands, or staples, with which it binds [10]. This approach allows the
staple strands to be added in excess concentration relative to the scaffold, which drives the self-assembly

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
4 https://doi.org/10.6028/jres.126.001
towards the desired structure. DNA origami systems may be designed to assemble into larger structures via
programmed interactions between multiple origami structures [11–13] (Fig. 3C).
While the fabrication of a new 3D structure can be readily accomplished without advanced background
knowledge, the design process itself is not intuitive. Fabrication of a structure by self-assembly converts
ssDNA strand sequence information into 3D topology via base pairing. However, the inverse operation
must be performed during the design process, i.e., converting a 3D structure into a linear sequence of bases.
While no individual step in the design flow is unusually difficult, an organized, iterative approach
significantly simplifies the process.
In attempting the design process, it is important to appreciate that current characterization capabilities
limit our understanding of the design-property relationships (like yield, mechanical strength, etc.). This
deficit in understanding increases the risk of failure for any one particular design and should be kept in
mind.
2. Basic Energetics, Yield, and Modeling of DNA Origami
While our understanding of the design-property relationships for many DNA nanotechnology systems
is insufficient to permit development and application of general, automated design tools (although
automated design tools that work within specific design constraints have been created [14–16]), there is a
consensus on many underpinning concepts, described here in brief. We begin with an examination of the
energetics driving the self-assembly of biological dsDNA, and we compare them with the energetics of
DNA origami. Next, we discuss the yield of the assembly process in creating the desired product. Finally,
we address common modeling and simulation tools used for these systems.
The energetics of classical dsDNA, i.e., Watson-Crick-Franklin base pairing, are predictable based on
the sequence and length of the DNA, where G-C pairings are more stable than A-T ones, resulting in a
higher melting temperature (T
m
) for G-C–rich sequences [17]. When dsDNA forms, the hydrophobic bases
are shielded from solution: Base-base interactions, called “base stacking,” along the long axis of the helix
are responsible for more of the favorable free energy of hybridization than the hydrogen bonding associated
with A-T and G-C base pairing.
Note: Future developments will doubtless improve our understanding of design-property trade-offs,
which are currently obscured by limited measurement capabilities.
While at this stage, many design choices are equivalent due to limited information, this document is
written in the hopes that future insights may be readily included into the design process described here.
Note: Notes will highlight considerations or pitfalls; red notes will indicate particular importance.
Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
5 https://doi.org/10.6028/jres.126.001
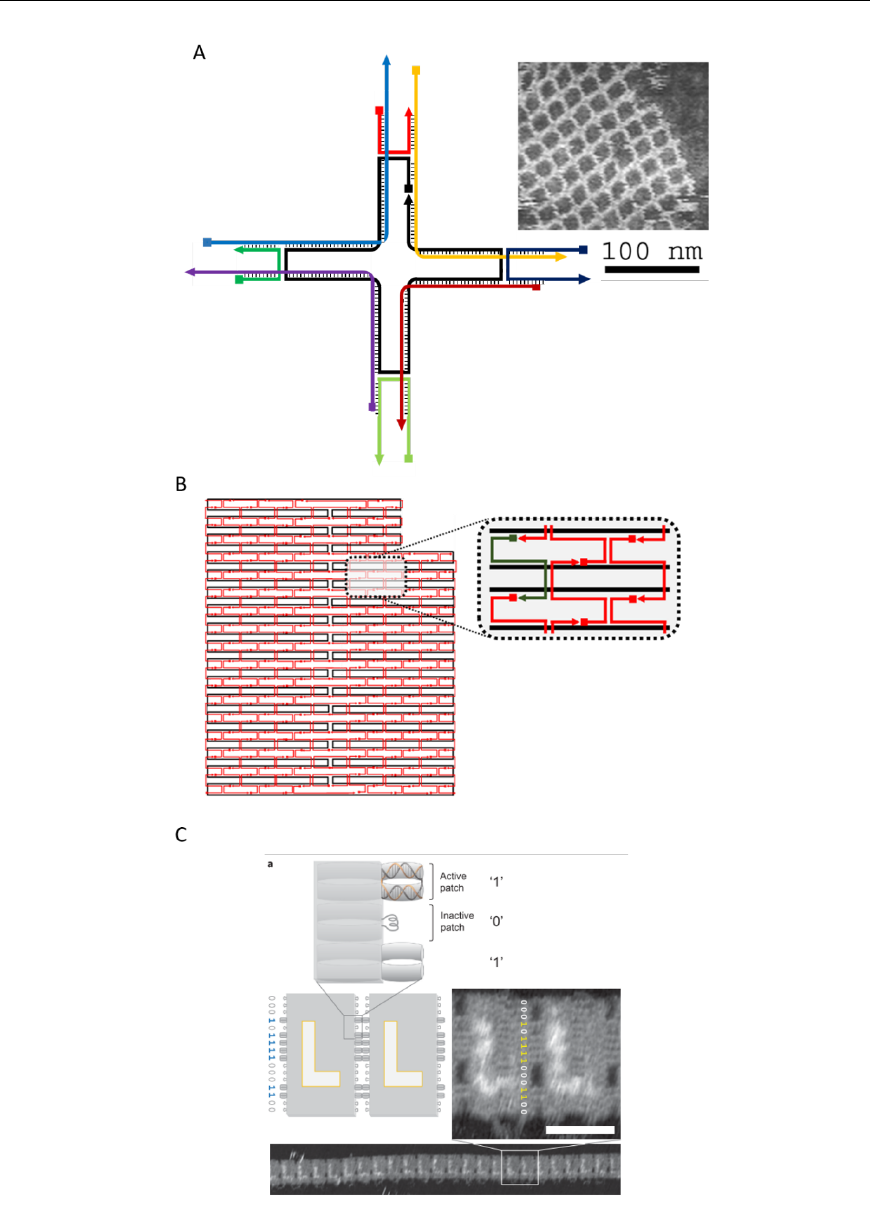
Fig. 3. Increasing complexity in assembly. (A) DNA tile, capable of 2D polymerization, comprising four crossovers [13]. (B) Typical
2D DNA origami, containing >400 crossovers. (C) Programmed stacking between origami structures [12].

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
6 https://doi.org/10.6028/jres.126.001
The energetics of DNA origami are at least an order of magnitude more complicated than classical
dsDNA [18]. Each ssDNA strand, or staple, in an origami has multiple subsequences, allowing it to form
one or more crossovers and jump between helices. Hybridization of these subsequences can behave
independently for topologically distant staples or cooperatively for nearby staples, where each “correct”
binding event energetically reinforces nearby correct binding events [18, 19]. This cooperativity
significantly improves the quality of assembly, but it also obscures our ability to measure and understand
assembly energetics. A first form of cooperativity occurs between subsequences on the same staple. When
a first subsequence on an ssDNA strand binds to the scaffold, the other subsequences on that particular
molecule have a dramatically increased probability of colliding with, and binding to, the scaffold. This can
be equivalently described using either a unimolecular equilibrium constant including an energetic penalty
for topological effects, or a bimolecular equilibrium constant with a j-factor that represents those effects
[20–22]. These topological effects are the second relevant form of cooperativity. There is an unfavorable
energetic contribution associated with conformational entropy loss between a free scaffold and one bound
into an origami. This penalty is universal across the structure, so an individual folding event will
cooperatively reduce the penalty for its neighbors. Finally, there is interstaple base stacking. If two
subsequences end immediately adjacent to one another along the scaffold, then the ending hydrophobic
bases, which would otherwise be exposed to water, are shielded from solution [23].
As with energetics, the yield of DNA origami is a comprehensive topic, and its full scope extends
beyond this document. Yield in these systems may be defined as the number of structures that have the
correct shape in microscopy, i.e., imaging yield, or it may be defined through the total number of discrete
staple binding defects, i.e., staple yield, or it may be defined through the ratio of appropriately incorporated
functional units to imaged origami, i.e., functional yield. While staple yield is the most difficult to measure
[24], it determines the other two, and it is the most relevant to design.
Staple yield defects, as illustrated
in Fig. 4, often take the form of missing or additional unprogrammed
staples. Individual defects are often too small to detect via imaging, and they may occur in locations
irrelevant for functional yield. In the illustrated case, the red/yellow/blue strands formed together first, but
instead of correctly binding the two subsequences on a single green strand, they bound either two separate
copies of the green strand or no copies of the green strand. The desired structure and the extra-strand defect
have similar energetics because the same sequences/lengths of dsDNA are formed, with the only difference
being a loss of translational entropy in the second copy. This loss is 3/2R, where R is the gas constant, and
T is the temperature. This malformed structure would then have two unsatisfied subsequences floating in
solution, which could potentially further polymerize. Generally speaking, these defects are avoided through
careful tuning of the relative concentrations of each strand, particularly for tile-based systems [6], and
through inherent cooperativity, particularly for origami [19].
Origami also has a natural stoichiometric control in which all staples bind to a single scaffold, allowing the
staples to be added in excess, and preventing missing-strand defects. For other systems, increasing the staple
concentration would dramatically increase extra-strand defects. However, the first form of cooperativity, described
above as the transition from bimolecular to unimolecular binding, favors the first staple molecule to bind to a
scaffold, preventing extra-strand defects even at high staple excesses. This is critical, because using high excesses of
staples allows one to drive the system away from missing-strand defects. In short, origami’s natural stoichiometric
control allows for dramatically higher yield of the desired structure for significantly less experimental effort. We
recommend the following papers for those interested in more depth on this topic [18, 25–28].
A related consideration is the synthesis yield of synthetic ssDNA. Phosphoramidite DNA synthesis
adds each base in a sequence with >99 % efficiency. However, the probability of synthesizing the correct
sequence in its entirety is this efficiency raised to the power of the number of bases. It is not uncommon for
a fraction of synthesized staple strands to be missing bases. If these missing bases occur on only one
subsequence, it can guarantee an extra-strand defect. The natural stoichiometric control in origami reduces
the likelihood of this, because a fully correct strand may bind in the second extra-strand position, where
cooperative energetic effects will favor it to exchange and displace the strand with the incorrect sequence.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
7 https://doi.org/10.6028/jres.126.001
Fig. 4. Examples of defect types for a primitive crossover.
While DNA origami assembly has been modeled and simulated, there are relatively few tools accessible
that do not require a significant investment to implement. However, simpler tools are available that provide
some useful insights. Web-based thermodynamics calculators predict classic DNA hybridization energetics
and predict unwanted hairpin formation [29–31], although these tools can typically only manage single
hybridization subsequences. CanDo is a finite element analysis tool, which treats the dsDNA helices as static
cylinders that can be torqued as they are welded together by crossovers and subject to thermal energy [32].
Finally, tools for molecular dynamics (MD), coarse graining, and multiscale modeling exist. MD simulates
behavior by treating atoms and bonds as beads and springs, respectively, with appropriate terms for
electrostatics, etc. [33–36]. MD is generally more accurate than coarse graining, which treats entire bases or
chemical subunits of bases as a single bead [37], but consumes so much more computational power that it
often cannot simulate events that occur at a timescale longer than nanoseconds. Multiscale modeling is a
technique by which different levels of coarse graining and MD are simulated and interchanged [38].
3. Common Terms
Scaffold
: Long DNA strand that is bound together to form an origami. It is often circular,
ssDNA, and sourced from a virus, although this is not always the case [39].
Scaffold routing: Raster pattern through which the scaffold travels back and forth through the structure.
Crossover: Position where DNA helices exchange ssDNA strands, linking them.
Staple: Short oligomer ssDNA sequences, usually synthetic, forcing the scaffold to route
into a desired shape.
Staple subsequence: Contiguous subset of the staple sequence that binds a contiguous sequence along
the scaffold. Subsequences are separated by crossovers.
Staple motif: Pattern of staple subsequence divisions that reoccurs throughout a structure, e.g.,
the 8 base-pair (bp) subsequence – 16 bp subsequence – 8 bp subsequence
commonly used in 2D origami such as the Rothemund Tall Rectangle [10].
Fold: The change in topology of the scaffold when a crossover, or ½ crossover, forms.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
8 https://doi.org/10.6028/jres.126.001
Sticky end: A short subsequence left intentionally unbound at a specific position in a
structure that will later be used to bind the structure and other nanoscale objects,
including nanoparticles, biomolecules, and other structures.
4. Instructions
Design methodologies are inherently unique because of system constraints. Due to the nature of DNA synthesis,
and specifically the minimum amount of material that it is feasible to synthesize or purchase, the time and money
needed to produce a prototype structure are approximately the same as those required for large-scale purchases. This
means that the number of practicable design iterations is limited by the expense of the design-prototype-test cycle.
This tutorial attempts to apply basic engineering design theory to minimize the number of design cycles.
As noted above, our ability to reduce the number of design cycles is hampered by our ignorance of
design-property relationships, which is, in turn, exacerbated by the limited characterization capabilities
currently available. To complicate matters further, the design space for DNA nanofabrication is large. In a
very real sense, our ignorance is a meaningful constraint; without knowledge of the design-property
relationships linking the items in Fig. 5, it is common for even experienced practitioners to get trapped in
an infinite design-iteration loop, searching for a “perfect” structure, which may or may not be physically
realizable. Additionally, given the limited number of design parameters that can be varied, any attempt to
optimize one property is highly likely to dramatically change the others.
Fig. 5. Examples of properties of interest, the attributes of a structure that determine them, and the relatively few design choices that
determine those attributes in a nonorthogonal fashion.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
9 https://doi.org/10.6028/jres.126.001
4.1 Predesign
As with almost any complex endeavor, the importance of predesign or preproduction cannot be
overstated. Given the relative immaturity of both the field of DNA nanotechnology and the nascent CAD
tool market created to serve it, the actual design process can be highly unintuitive. Beginning design
without a meaningful list of specifications invites significant loss of time.
Defining Functional Requirements:
a. List all physical features relevant to the purpose/application, such as level of control needed over
distance between specific functional components, overall footprint, accessibility of components to
solution, chirality, stimuli response, dynamic motion, inter-structure interactions, etc.
b. Determine how success in creating the desired structure will be determined.
b.I. During development:
for example, transmission electron microscope (TEM), atomic force microscopy (AFM), gel
electrophoresis.
b.II. During use:
for example, pharmaceutical activity, total number of actuation cycles before failure.
c. Order these physical features by their impact on the probability of success in application.
c.I. Combine the above information into a single set of specifications.
c.II. Add fabrication constraints, e.g., monetary constraints on number of prototyping cycles.
Two constraints that pertain to the majority of origami are the length and, commonly, the circularity of
their scaffolds, although both can be circumvented through application of production and enzymatic
methods [40]. Practically, this means that most structures are limited to a maximum of 7 249 bases,
corresponding to a dsDNA length of 2 389 nm or a surface area of approximately 5 000 nm
2
(6 000 nm
2
for
2D structures after relaxation and expansion).
The ubiquity of the most common scaffold, the M13mp18 genome, is in part historical and in part
practical. The practical considerations are as follows. The M13 bacteriophage infects the bacterium
Escherichia coli (E. coli), both of which have been researched exhaustively. The M13 bacteriophage is well
situated in inherent trade-offs among the type of virus (ssDNA, dsDNA, ssRNA, dsRNA), the length of the
viral genome, and the mutation rate [41]. While the M13 bacteriophage packages its genome as ssDNA,
which is highly desirable for origami, its packaging accommodates varying lengths of ssDNA without
significant mutation concerns. The cylindrical viral capsid consists of a handful of proteins at the caps and
a repeating coat protein along the cylinder wall. For the M13 phage, changing the amount of ssDNA
packaged can be accommodated with a change in the number of sidewall proteins and the concomitant
length of the viral capsid cylinder. The use of the mp18 sequence specifically, rather than another version
of M13, is largely historical, since it was used in Paul Rothemund’s initial work on DNA origami [10].
While longer and shorter scaffolds have been developed and employed [42, 43], the success and utility of
the original have limited the drive to explore this design variable in detail.
4.2 Macroscale Prototyping
Because true nanoscale prototypes are infeasibly expensive in both time and money, macroscale
prototyping is critical to efficient design. Mockups can be made with pen and paper, dowel rods, magnetic
toys, etc. Note: Colored magnetic bars are particularly useful for this purpose.
Note: Given our current knowledge, subjective choices are unavoidable.
Caveat Lector: Hic Sunt Dracones.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
10 https://doi.org/10.6028/jres.126.001
Creating mockups is largely iterative, as final and initial designs are rarely identical. If at any point it is
likely that the specifications and constraints cannot be satisfied for a given structure, reiterate from the most
relevant step.
a. Build a mockup (Fig. 6).
Attributes of effective mockups include the following:
• It is easy to label and relabel subsections.
• They are easy to assemble/disassemble.
• They have rotatable joints for easier flattening.
b. Estimate whether specification is feasible.
For example, for each edge/surface of a structure, determine:
• How many helices will run through this edge/surface?
• Given the number of helices and relative proportions, can the physical specifications and the
constraint on dsDNA length be met simultaneously?
Fig. 6. Tetrahedron mockup.
c. “Flatten” the mockup (Fig. 7).
Both structure design by hand and most available CAD tools route the scaffold in 2D and force all
helices to appear as if they are parallel. It is therefore important to be able to flatten the model to
visualize the relationship between the design and the CAD representation.
Fig. 7. (Left) 3D mockup. (Right) Flattened mockup.
Note: Only check as many specifications/constraints as are relevant at each step.
Many specifications can only be estimated later when the design is more fully realized.
If specifications/constraints cannot be satisfied, try a new shape.
This step will be repeated throughout the design process.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
11 https://doi.org/10.6028/jres.126.001
d. Identify and label the subsections and their interconnections (Fig. 8). For an additional example of
the importance of this concept, see CAD drawing in Fig. 12.
Fig. 8. (Left) Annotated, flattened mockup schematic including topological links. (Right) Flattened mockup schematic after
parallelization of helices.
e. Select the helix arrangement, also called a scaffold lattice (Fig. 9), and mock up a routing pattern (Fig. 10).
2D
3D
Scaffold
Lattice
or
Helix
Arrangement
Fig. 9. Scaffold lattice examples for honeycomb (top) and square (bottom) lattices. Dotted circles indicate potential locations for other
helices, and full cylinders indicate helices in the design.
Note:
Lattice choice will constrain later options for crossover distances/density. However, it is preferable to
make only one design choice at a time and iterate, so it is advisable to focus on accommodating the
circular nature of the scaffold and length/size constraints at this stage.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
12 https://doi.org/10.6028/jres.126.001
Fig. 10. Physical mockup of scaffold routing using a beaded chain.
f. Using the mockup routing pattern, draw a full routing pattern (Fig. 11).
2D 3D
Scaffold Routing
Fig. 11. Example routing patterns for (left) 2D square lattice structures and (right) 3D hexagonal lattice structures.
Note:
Square lattices easily lie flat on surfaces but have difficult-to-manage internal strain (Sec. 4.5).
As a default, the square lattice is preferable for surface applications or if AFM will be the primary
imaging technique. The hexagonal lattice is typically better suited to 3D applications.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
13 https://doi.org/10.6028/jres.126.001
g. Repeat step b—As the rough design is now more complete, confirm that specifications can be met:
• Is it physically possible to perform a valid scaffold routing given that the scaffold is circular?
• Are length/size specifications still feasible?
4.3 Computer-Aided Design (CAD)
For convenience, the cadnano2.0 [1] (cadnano.org/ & github.com/douglaslab/cadnano2) design tool
will be used as an example in this section. Other CAD tools, such as scadnano (scadnano.org/), SAMSON-
adenita toolkit [44] (www.samson-connect.net/elements.html), vHelix [45] (vhelix.net/), and Daedalus [46]
(daedalus-dna-origami.org/), are readily available and may have their own advantages. As many of these
tools are open-source tools, we used a GIT tool, GitHub desktop, to ensure operation of the most recent
version and to minimize installation problems.
Figure 12 shows two example designs and CAD screenshots. The visual complexity associated with
the routed scaffold and 200+ interconnected staples can make CAD modification difficult. Because of this,
emphasis is placed on carefully planning both the design and the CAD file to make both visualization and
modification less cumbersome. Some CAD tools allow for rotation of helices within a design (Fig. 12),
which can reduce some, but not all, visual clutter.
Note: This tutorial will only address the software aspects of initial design steps. The reader is advised
to consult user documents for their CAD tool of choice.
Note: It is impossible to avoid significant visual clutter when representing the interconnections present
in even a simple 3D structure.
Because available CAD tools can be inconvenient for arbitrary modification of designs, a minimally
cluttered design document streamlines new structure development.
The right side of Fig. 12 provides an example in which visual clutter could become problematic. If
empty helices were not placed between bundles A and F, or if the vertical placement of the bundles was
mismanaged, it would be difficult to identify individual staple locations if, for example, one wished to
label the midpoint of two bundles with unique nanoparticles.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
14 https://doi.org/10.6028/jres.126.001
Notched Rectangle ( 2D )
Tetrahedron ( 3D )
Fig. 12. Examples of origami design and CAD screenshot. (Left) Notched rectangle structure. (Left-Top) Graphic representation of
design. (Left-Bottom) Screenshot of CAD screenshot. (Right) Tetrahedron structure. (Right-Middle) cadnano CAD screenshot.
(Right-Bottom) scadnano CAD screenshot.
h. Define the workspace (Fig. 13A–D).
h.I. Select helix type (Fig. 13A).
h.II. Define helices (Fig. 13B–C). For cadnano, the left window looks down the z axis of the helices,
while the right window has the z axis running from left to right.
h.III. When helices on the left are clicked, an “empty” helix on the right will appear.
In cadnano2.0, the helices cannot be reordered. It is advisable to add “empty” helices between
substructure units to make them visibly distinct from one another, e.g., A–F in the tetrahedron
in Fig. 12.
h.IV. Clicking again, holding, and dragging across the helices on the left will automatically generate
a scaffold routing between all the helices over which the mouse was dragged over (Fig 13D).
Note: This step must often be reiterated to ensure the CAD workspace is easily legible.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
15 https://doi.org/10.6028/jres.126.001
A
B
C
D
Fig. 13. (A-D) CAD images corresponding to step h. (A) Lattice selection determining staking of helices when looking down the z
axis. (B) Selection of helices to populate into CAD file. (C) Populated helices for a six-helix bundle. (D) Automatic scaffold routing
created by dragging mouse across helices in C.
Note: cadnano v2.0 centers the new routing around the yellow slider in the right window.
Note: Any CAD tool will automatically track potential crossover spacings for both the scaffold and
staples given the properties of B-DNA and the positions of neighboring helices.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
16 https://doi.org/10.6028/jres.126.001
i. Test staple motifs on a small section of the scaffold routing pattern. For examples, see Fig. 14.
i.I. The goal, at this step, is to confirm that a valid staple motif exists for the chosen scaffold routing
that can meet the design specifications and constraints.
i.II. Staples may be drawn manually, or they may be edited after generation via cadnano’s
“autostaple” function.
o Consider the following criteria when evaluating staple motifs and design specifications.
Consider the staple fit over the scaffold “seams,” because positions where the scaffold crosses
over between helices are typically the longest distance folds, and the most susceptible to
defect formation:
o Fewer than 5–8 bases of connection may not have desired stability.
Consider the distance between, and number of, crossovers:
o More crossovers => stiffer structure (along crossover axis).
o More crossovers => more internal strain (addressed in initial testing and fine tuning).
Consider the 5′ and 3′ (start and stop) position locations that are natural locations to extend
sticky ends and create functional binding sites (addressed in additional conceptual tools
section).
Note: cadnano v2.0 indicates the rotational position of the scaffold bases at each helix underneath the
yellow slider as a blue line on the helix in the left window, Fig. 13D).
The blue line indicates the position of the minor groove, where strands exit/enter the helix. A more
general description is given in Sec.5.3.
Visual Clutter Management Checklist
In current software versions, the helix positions on the right window cannot be modified. Care should
be taken with their arrangement to minimize visual clutter.
We advise the following approach.
Visual proximity on the right window schematic is dependent on the order in which the helices are
activated, step h, on the left window. To minimize visual clutter, the helices should be activated in an
order that groups substructures.
1. Identify the number of substructure sections in the mockup.
a. Notched Rectangle = 1 substructure (Fig. 12).
b. Tetrahedron = 6 substructures (Fig. 12).
c. Tower = 5 substructures (Fig. 18).
2. For each substructure, count necessary helices and identify helices/positions where strands
will jump to other substructures (Fig. 8).
3. Label mockup substructures in an order that shortens intersubstructure connections in
flattened/parallel view.
4. Label helix order within substructures to place intersubstructure connections at the top/bottom
of that substructure in the flattened/parallel view.
5. Starting with the first substructure, activate helices in order.
6. Between substructures, activate at least two helices to provide visual space between them
7. Click/fill helices, step h.III, in the same order as above.
a. If the automatically filled scaffold is unsatisfactory, then it is advisable to undo the
action, move the yellow slider that centers the automatic filling function, and try
again before resorting to manual drawing tools.
8. SAVE all intermediate drafts as separate files.
Note: It is not possible to represent a 3D structure as a flattened and parallel line schematic without
crossovers running “over” the rest of the design. While no arrangement will prevent problematic visual
clutter, CAD workspace organization can significantly reduce it.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
17 https://doi.org/10.6028/jres.126.001
Fig. 14. Example staple motifs on the square 2D lattice, with tentative naming convention based on the number of bases in each
subsequence with differentiation for full and half crossovers where necessary.
j. For testing, we advise the following steps (Fig. 15).
j.I. Confirm the autostaple pattern has satisfactory crossover options; if not, then delete and recenter
the scaffold routing (Fig. 15A).
j.II. Treat the scaffold seam regions separately from the rest of the structure.
j.III. Identify a staple motif rule, e.g., one 8 base subsequence, one 16 base subsequence, and one 8
base subsequence, preferably with a nucleating subsequence [47].
k. Save a copy of the scaffold routing empty of staples (Fig. 16).
l. Attempt to tile selected motif over a small area. Write down or otherwise confirm the tentative pattern
of the staple motif.
m. Estimate feasibility of meeting specifications in step b.
n. Populate the routing pattern with the tentative staple motif choice from step j.III. An example of this is
shown in Fig. 12.
o. Save the completed file under a new file name.
Note: Synthetic ssDNA synthesis is typically limited to <60 to 100 bases.
The yield of a synthetic ssDNA is roughly equal to 0.95
N
to 0.99
N
, where N is the number of bases.
Long staples have lower yield but theoretically allow higher thermal stability because longer DNA
domains melt at higher temperatures.
Note: Staple motifs, particularly for multilayer 3D objects, are one of the few choices for which there is
a known failure mode. Staple motifs consisting only of equal-length subsequences have a failure in 3D,
honeycomb lattice structures and have been known to result in failed annealing and aggregation.
Where possible, staples should be designed with a “nucleating” subsequence of longer length [47].

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
18 https://doi.org/10.6028/jres.126.001
A
C
B
Fig. 15. Example tests for staple motifs. (A) Initial scaffold routing. (B) Staple crossover pattern generated by the autostaple function.
(C) Incremental division of the staple crossover pattern into individual staples of a length that can be ordered for custom synthesis.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
19 https://doi.org/10.6028/jres.126.001
Notched Rectangle ( 2D )
Tetrahedron ( 3D )
Fig. 16. Example routing maps and final staple screenshots matching Fig. 12.
Note: Step n often reveals flaws in the staple motif choice or scaffold routing. Expect one or two
reiterations at this stage.
Create backup files for each staple motif and routing choice.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
20 https://doi.org/10.6028/jres.126.001
4.4 Initial Structure Prediction
Numerous tools exist for evaluation of structures. While we use CanDo (https://cando-dna-
origami.org/), other tools, such as circleMap (https://nanohub.org/resources/cadnanovis) and University of
Illinois at Urbana–Champaign (UIUC) multiscale (http://bionano.physics.illinois.edu/origami-structure),
also exist. Physics-based simulations such as UIUC multiscale or oxDNA [37] may provide more nuanced
predictions at the cost of time and expertise to initiate. An advisable first step is to test new structures with
a finite element analysis tool such as CanDo. Finite element analysis treats each helix as an independent
cylinder and the crossovers as welds with nominally realistic levels of torque and strain given the angle of
the crossover and physical properties of DNA. Finite element analysis provides a preliminary confirmation
of the 3D structure, and it is preferable to performing an MD, coarse-grained, or multiscale modeling
simulation on the structure, which could potentially waste significant computational resources.
If curvature of the structure is irrelevant to the design specifications, then the need for testing and fine
tuning may be limited. Figure 17 shows the CanDo simulations on the notched rectangle and an AFM
image thereof. The curvature in this CanDo simulation is a reasonable representation of the effects of
internal strain and torque on 2D origami structures [48]. A 3D tower structure on the square lattice is shown
in Fig. 18.
For CanDo, red indicates a section of the structure that is freer to move, while blue indicates a static
section. CAD modification at this stage is highly dependent on design specifications. One common
modification, strain compensation for square lattice structures, will be addressed in the next section.
Fig. 17. Notched rectangle design, CanDo result, and AFM image.
Note: Finite element analysis is sufficient to predict general shape and requires minimal computing
resources. If it is necessary to perform MD or multiscale modeling for a more detailed representation
of the structure, it is advisable to wait until after the user is satisfied with the structure’s performance
in finite element analysis modeling.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
21 https://doi.org/10.6028/jres.126.001
Fig. 18. Tower origami CanDo result.
4.5 Crossovers and Internal Strain
When addressing internal stress, strain, and torque of DNA origami, it is doubly important to know in
advance which functional requirements one is pursuing. While the interplay among design, stress, strain,
and torque is relatively easy to model, they are often difficult to visualize and implement. As a result, it is
easy to waste significant time and effort modifying these factors to no positive effect.
Internal strain and torque in DNA origami are well described by the analogy of welding together
springs, as illustrated in Fig. 19, where the metal of the spring represents the minor groove. In this analogy,
there are two springs that must be joined at two positions to create a larger structure. Both springs have the
same helicity and period. After the first weld, the springs are aligned, and there is a visible periodicity
where the metal of each spring touches. At these positions, one could apply a second weld with no tension
or torque. However, if one wished to weld at two positions of equal distance to the first weld, but a little
before or after this period, one would have to tighten or loosen the spring to bring those points into line.
Similarly, if one wished to weld the springs at slightly different lengths, one would simultaneously have to
compress and tighten one spring while stretching and unwinding the other.
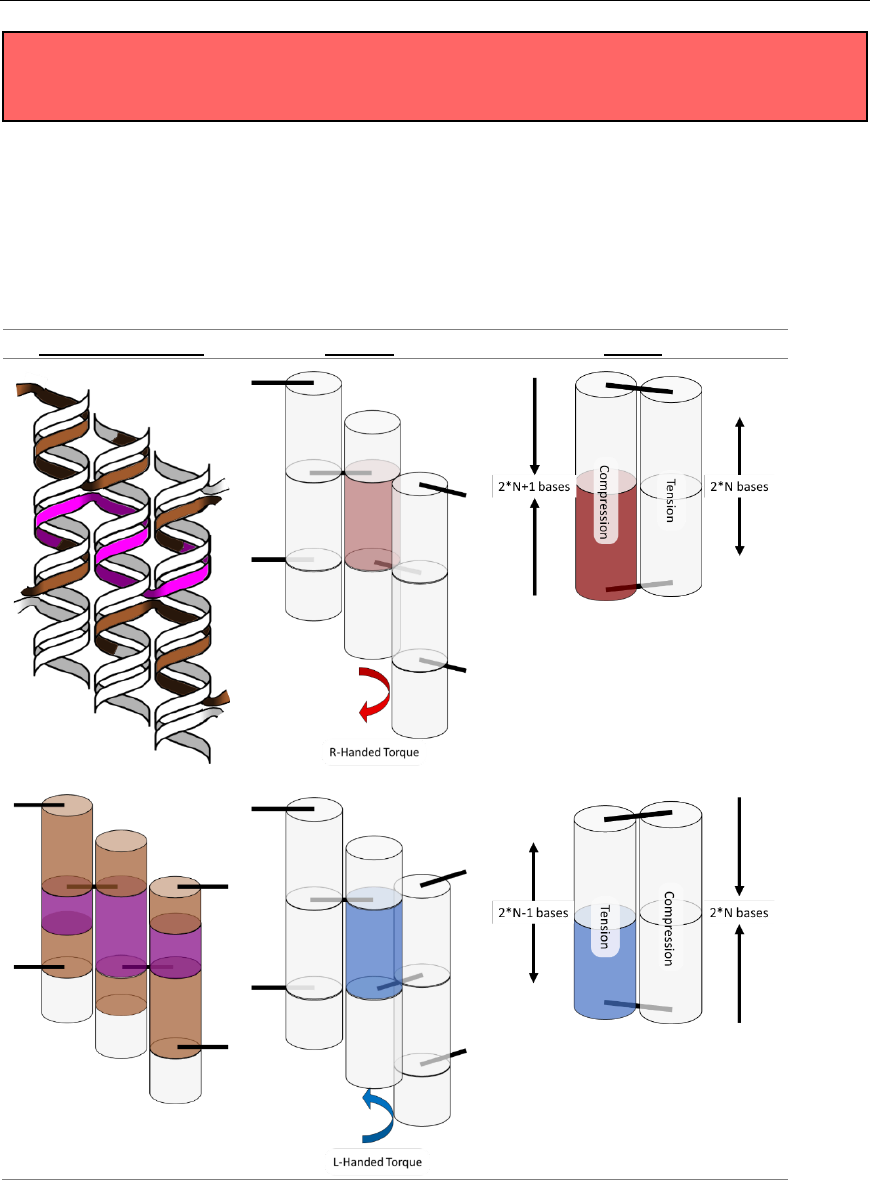
Fig. 19. Spring analogy for over- and underwinding of DNA when helices are connected by crossovers. L-handed and R-handed
indicate left-handed and right-handed, respectively.
Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
22 https://doi.org/10.6028/jres.126.001
The spring analogy is imperfect, DNA cannot be treated as continuous, and crossovers can only occur
between integer numbers of bases. This can be problematic because the dsDNA helix has 10.5 bases per
full turn. The number of bases between potentially relaxed crossover positions depends on the angle
between helices being welded (Fig. 23), which can confuse the process of choosing the number of bases
between crossovers for a particular design. Binding numerous helices at a variety of angles is much more
complex than simply welding two springs.
Relaxed/Example
Torque
Strain
Fig. 20. (Left) Cylinder representation of dsDNA between crossovers. (Middle) Resulting torque within a structure if only the center
purple helix/cylinder from the left side is overwound or underwound. (Right) Schematic of strain in DNA crossovers. Torque and
strain will occur simultaneously in most systems as the sum of small contributions from all crossovers. They are depicted here as
independent for illustrative purposes only. As in Fig. 19, red indicates overwound helices, and blue indicates underwound helices.
Note: Modification/control of torque and strain in DNA nanostructures can be very time-consuming.
Before attempting to modify a structure, take time to confirm that it is necessary for your functional
specifications.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
23 https://doi.org/10.6028/jres.126.001
The torques and compressive/tensile forces may be controlled by varying the number of bases between
crossovers (in cadnano v2.0, by using the skip and insert tools). The effects of these approaches are shown
schematically in Fig. 20 and Fig. 21. Additionally, Fig. 22 uses a full origami CAD example.
The constraints on staple length dictated by yield in ssDNA synthesis and specifications for flexibility
(requiring more/fewer crossovers) are important to consider because torque and strain are controlled via
changing crossover spacing with skips and inserts. The default spacing is a set number of bases in the
software for each lattice/angle between helices. This number must be an integer, but the relaxed number of
bases for that angle is often not an integer. A skip, when placed between crossovers, informs the software
to have one base fewer than the default between those crossovers. An insert informs the software to have
one base more than the default.
Fig. 21. (Left) Arrangement of strained/torqued helices. (Right) Approximate effect on overall structure conformation of that
strain/torque arrangement. As in Fig. 19, red indicates overwound helices, and blue indicates underwound helices.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
24 https://doi.org/10.6028/jres.126.001
Twist
Schematic
Average
Twist
Values
Default Length:
slightly overwound
10.66 bp/turn
33.75°/bp
Alternating Skips:
10.333 bp/turn
34.84°/bp
¼ Skips:
10.5 bp/turn
34.28°/bp
¼ Skips staggered:
10.5 bp/turn
34.28°/bp
CanDo
Prediction
Fig. 22. CanDo example of DNA skips and strain compensation on a square lattice for a planar origami. The top row shows the
handedness of the torque as determined by the presence and location of skips. Red indicates right-handed torque helices, and blue
indicates left-handed torque in the twist schematic. The average twist values are indicated for each of the four designs. As noted in the
text, the number of base pairs per turn in an unstrained dsDNA helix is 10.5. The bottom row shows the outputs of the corresponding
CanDo models. These illustrate the approximate shape and local helix mobility.
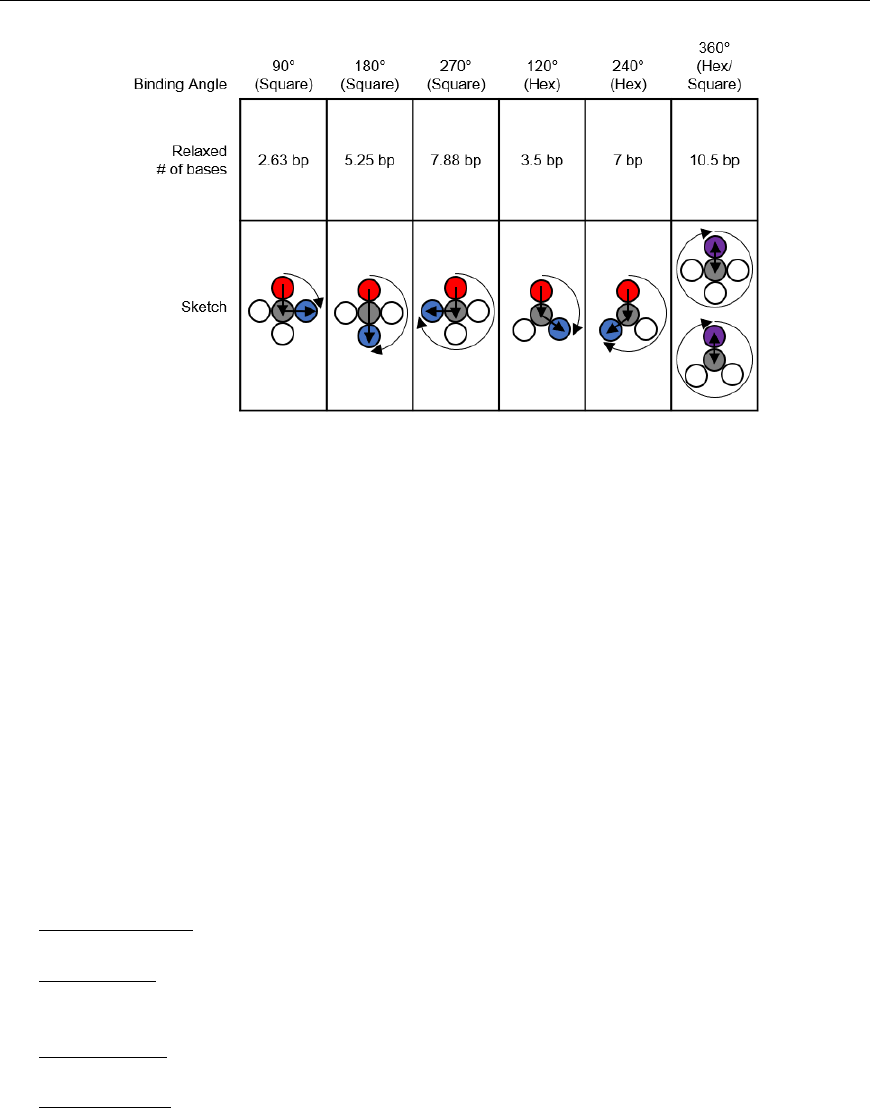
The “relaxed” number of bases between crossovers depends on the enforced angle between helices, as
shown in Fig. 23. While this may be a useful reference, it is a worthwhile exercise to independently derive
these numbers. On the hexagonal lattice, the relaxed number is either an integer or ± 0.5 bases. While the
square lattice lies flat for easier AFM imaging, compensation of residual strain/torque is much easier for the
hexagonal lattice.
The reader is advised to save backup copies of the CAD file between modification attempts, and to
regularly use finite element analysis prediction for each step to ensure specifications can be met. Given the
number of files generated, a rational file naming and organizing scheme is essential.
Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
25 https://doi.org/10.6028/jres.126.001
Fig. 23. Relaxed number of bases for common helix binding angles. If the distance between two crossovers is greater than a full turn,
to determine whether it is over- or underwound, subtract the largest integer multiple of 10.5 bases possible without resulting in a
negative number.
Note: Use of this strain to generate curved surfaces is addressed in Ref. [49].
5. Additional Conceptual Tools
5.1 Folding Information and its Representation
A common concern for designers of DNA origami systems is, among the multiplicity of design
options, how to determine if one is better or worse than others. Current understanding is insufficient to
answer this question, but a few conceptual tools can illustrate how structures are different.
One factor that is known to be important in origami folding are entropic penalties, which occur when
the topology of the scaffold is forced to change via staple binding. The total entropic cost of transforming
the flexible ssDNA scaffold into a single relatively inflexible block is small compared to the energy gains
associated with binding. However, this cost is levied unevenly, and it significantly discourages long-
distance folds from occurring until other staples reduce that distance. Fold distances may be visualized
within the routing pattern, circle plots, histograms, and square plots.
The routing pattern
, discussed in the instructions, conveys the desired shape of a structure and is a
central focus of the design process.
The circle plot illustrates the initial distance each fold must bridge. In the circle plot, the scaffold is
represented by the outside circle, while each arc indicates a fold. One limitation of the circle plot is that the
vast majority of folds are too small to see clearly.
Histogram plots of fold distances readily depict small folds and can be used to determine fold-distance
statistics, but they contain no information on fold nesting.
The square plots that we have devised, and that we present here for the first time, attempt to combine
the most useful aspects of circle plots and fold histograms. The scaffold in the square plot is represented by
the x axis, and in the case of a circular scaffold, it should be noted that that origin and the furthest extent
represent adjacent points on that scaffold. Folds are plotted as lines between their start/stop positions, with
the y-axis position corresponding to the initial fold distance, equal to the difference between the start and
stop positions on the scaffold for that fold. Folds plotted above/below each other will directly reduce the
looping entropy penalties for each other. In this way, cones that run from the top to bottom of the square
plot indicate folding pathways along which a long stretch of scaffold is “zipped” closed. To emphasize this

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
26 https://doi.org/10.6028/jres.126.001
feature, a transparent shading was applied from each fold down to the bottom of the plot. Darker regions
reflect many nested folds on top of one another. In short, darker shading regions imply a folding “pathway”
in which some shorter folds must occur first to enable subsequent folds.
5.2 Congruent Shapes of Various Scaffold Routing
To illustrate the plots described in Sec. 5.1 and the plethora of design choices, we created four different
DNA origami of identical, or nearly identical, 2D footprint. In an AFM or TEM, these structures would be
difficult to distinguish from one another.
The two designs most typical of 2D origami to date are the vertical and horizontal designs (Fig. 24).
The scaffold routing for these two designs places a seam that bisects the rectangular footprint along a
cardinal axis. As the rectangle has a high aspect ratio, this results in very different loop distributions. The
vertical origami has a very large number of short-fold-distance nucleating sites, but many more long-
distance folds as the system zips up the seam. In contrast, the horizontal origami has relatively few very
long or very short folds, and a large number of medium-distance folds.
The other two designs presented here are less typical but represent conceptually simple modifications
to the horizontal design. The horizontal interdigitated design moves the scaffold crossovers of the seam
back and forth so that they are no longer contiguous, broadening the distribution of fold distances. The
woven structure is the result of taking each crossover position and, if possible, having the scaffold jump
between helices rather than the staple.
In evaluating a design, we recommend use of the circle plot to confirm the number and symmetry of
substructural units in the origami, as the circle plot can be visualized on “top” of the routing for
troubleshooting. A drawback of the circle plot, shown in Fig. 25, is that it poorly depicts the very short
folds, which often comprise the majority of a structure. The square plot is more useful for identifying the
degree to which folds will occur in parallel or via a few path-dependent “zippering” interactions.
The examples in Fig. 24 illustrate design differences in cases where numerous designs occupy the
same footprint. Note: The routing patterns, particularly for the vertical origami, are not to scale.
The “horizontal” origami has helices running parallel to its long side and a central seam, and it would
be rigid in the long direction but flexible along the short direction. If one only looked at the routing map, it
would be easy to believe this origami has the greatest number of long folds because the scaffold has the
fewest turns.
The “vertical” origami has helices running parallel to its short side and a long central seam, and it
would be flexible in the long direction but rigid in the short direction. Because there are many positions
where the scaffold routes back and forth in the short direction, there are large numbers of short folds,
shown in the histogram.
The “interdigitated horizontal” origami illustrates how the fold distances in the horizontal origami
would change if the seams were shifted/interdigitated. This example shows how simple changes to the
routing pattern can dramatically change the distribution of loop distances.
Finally, the “woven” origami takes the horizontal origami design and maximizes the number of
positions where the scaffold is jumping between helices, rather than the staples.
While different routing patterns result in quite different fold distance distributions, we do not yet know
if those differences have a significant impact on the quality of self-assembly.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
27 https://doi.org/10.6028/jres.126.001
Fig. 24. Example representation of congruent designs. For each design, Top-Left: scaffold routing, Top-Right: histogram, Bottom-
Left: circle plot, Bottom-Right: square plot.
Horizontal Origami
Horizontal Origami Interdigitated
0
50
100
0 1000 2000 3000
0
50
100
150
Counts
Fold Distance
Cumulative Percent
0
50
100
0 1000 2000 3000
0
50
100
150
Counts
Fold Distance
Cumulative Percent

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
28 https://doi.org/10.6028/jres.126.001
Fig. 24. Example representation of congruent designs. For each design, Top-Left: scaffold routing, Top-Right: histogram, Bottom-
Left: circle plot, Bottom-Right: square plot (continued).
Woven Horizontal Origami
Vertical Origami
0
50
100
0 1000 2000 3000
0
50
100
150
Counts
Fold Distance
Cumulative Percent
0
50
100
0 1000 2000 3000
0
50
100
150
Counts
Fold Distance
Cumulative Percent

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
29 https://doi.org/10.6028/jres.126.001
Fig 25. Zoom-in of respective circle plots illustrating the short folds, which can be difficult to see in circle plots.
5.3 Staple Extensions for Addressing Functional Units
Sticky ends—short lengths of unbound ssDNA—are used to create addressable sites on the origami to
which functional units such as nanoparticles or biomolecules may be added. These sticky ends may be
incorporated as a hairpin in the center of a staple or as an extension at either end of the staple. In both
cases, it is critical to know which direction relative to the bulk of the origami the sticky end will exit.
As briefly mentioned in the introductory material, the extension will exit the helix based on the
position of the minor groove at the relevant base position (Fig. 1 and Fig. 2).
This process is illustrated in Fig. 26. The simplest visualization comes from combining the direction of
the staple, the direction of the nearest crossover, and the helicity of dsDNA to determine the exit direction for
the sticky end via the right-hand rule. This can be done by imaging the right-hand-thumb in the 5′→ 3′
direction and the index finger initially in the direction of the crossover. As the hand moves along the dsDNA,
the fingers should curl at ≈ 34.3°/base in the 3′ direction. This is approximately 270° per every 8 bases.
Fig. 26. Design example and application of the right-hand rule in placing an addressable sticky end.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
30 https://doi.org/10.6028/jres.126.001
One useful exercise is to examine the Rothemund Tall Rectangle [10], which is the basis structure for
the notched rectangle shown throughout this text. The Tall Rectangle is often used as a representative 2D
structure. Based on its design, all its 5′ and 3′ staple ends exit the origami on the same face. If functional
components are needed on both sides of the structure, a minor redesign of the staples is necessary.
Confirming this property, and attempting said staple redesigns, is a worthwhile exercise for designers
new to DNA origami.
5.4 Programmed and Unintentional Intra-origami Base Stacking
In the introduction, origami were described as being built from dsDNA helices welded together by
crossovers. There is an additional critical feature of these helices relevant to DNA origami. While the radial
surface of the helix comprises the negatively charged hydrophilic backbone, the end caps comprise the
hydrophobic bases. The sides of an origami parallel to the helix direction are hydrophilic, while those
perpendicular to the helix direction (with exposed cylinder ends) are hydrophobic.
As such, the exposed cylinder ends at the edges of DNA nanostructures are prone to reversible, low-
energy stacking [12], as shown in Fig. 27.
This stacking can be beneficial, e.g., for surface tiling, or disadvantageous, because it may lead to
uncontrolled agglomeration. A tried-and-tested of way to prevent stacking is to place a 4+ base poly-T loop
on the staples when they jump between helices at the cylinder ends. If G-C hairpins are placed in these
positions on the staples, they can create physical lock-and-key arrangements that only allow stacking
between preprogrammed substructures.
Fig. 27. Interorigami base stacking between helix ends, prevention thereof, and programmed base stacking.
5.5 Ordering and Naming Conventions
Most DNA nanofabrication CAD tools have a primary export format of comma-separated value, or
.csv, files. This is because most oligonucleotide synthesis companies perform bulk orders through .csv

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
31 https://doi.org/10.6028/jres.126.001
workbook files. While the relevant cells from the .csv file can be copied directly into the order forms, it is
advisable to take additional time to consider naming conventions.
One DNA origami typically requires 250 staple strands or more, requiring procurement of three 96
deep-well plates, with numerous individual tubes for functionalized staples. Given that one set of plates can
last for both a considerable time and for a considerable number of experiments, a consistent naming
convention is useful.
CAD tool .csv export files will typically include information such as the helix number and base
position along that helix for each staple. It is advisable to use this information as part of the name in
purchasing, to minimize frustration in future attempts to add or remove individual staples.
Toward this end, common text extraction and concatenation tools in workbook programs are useful.
Examples include LEFT() and “&” in Excel
®
. Combining these with an autofill function can allow for
complex names to be quickly generated for all strands. It is advisable to include properties like a structure
name abbreviation, substructure, and helix/base position of the staple.
Default mass ordering spreadsheets will often name the plates “Sheet1” to “SheetN” or “Plate1” to
“PlateN.” The wise researcher will avoid a freezer full of many structures each with multiple plates labeled
with such generic names.
6. Summary
By breaking the design process for DNA origami into a sequence of iterative operations involving
straightforward decisions and prioritizations, it is possible to streamline the creation of a new design, both
minimizing nonproductive tinkering and reducing the likelihood of errors. The general flow of this process
is applicable to other types of DNA nanofabrication systems such as single-strand tiles (SSTs) and
infinitely tiling systems.
If only one piece of advice from this document is retained by the reader, we strongly suggest that it be
the understanding that critical and intentional predesign, including the generation of a complete list of
functional requirements, is absolutely necessary to the efficient development of a working structure.
Acknowledgments
The authors would like to acknowledge Nicole Estrich for providing the 3D tetrahedron origami as
well as for useful discussions on the design process.
Contact Information
Jacob Majikes, research scientist, Microsystems and Nanotechnology Division, Physical Measurement
Laboratory, NIST, Jacob.majikes@nist.gov.
Note: As one set of origami plates can last for an extended period, it is reasonable to assume
that another researcher will have to use your notation to understand the structure.
Pay it forward by using a clear and consistent naming convention for the plates & strands.
Note: Using the helix number and base position, or other identifying feature in your CAD tool
of choice, in a naming convention will allow you or your successor to easily find and replace
individual staples for subsequent experiments/sticky strand labeling.

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
32 https://doi.org/10.6028/jres.126.001
7. References
[1] Douglas SM, Marblestone AH, Teerapittayanon S, Vazquez A, Church GM, Shih WM (2009) Rapid prototyping of 3D
DNA-origami shapes with caDNAno. Nucleic Acids Research 37(15):5001–5006.
[2] Castro CE, Kilchherr F, Kim D, Shiao EL, Wauer T, Wortmann P, Bathe M, Dietz H (2011) A primer to scaffolded DNA
origami. Nature Methods 8(3):221–229. https://doi.org/10.1038/NMETH.1570
[3] Kim D, Kilchherr F, Dietz H, Bathe M (2011) Quantitative prediction of 3D solution shape and flexibility of nucleic acid
nanostructures. Nucleic Acids Research 1(7):1–7. https://doi.org/10.1093/nar/gkr1173
[4] Carlson R (2009) The changing economics of DNA synthesis. Nature Biotechnology 27(12):1091–4.
https://doi.org/10.1038/nbt1209-1091
[5] Bloomfield VA, Crothers DM, Tinoco I (2000) Nucleic Acids: Structures, Properties, and Functions ed Stiefel J (University
Science Books, Sausalito, CA 94965).
[6] Seeman NC (2016) Structural DNA Nanotechnology (Cambridge University Press, Cambridge).
https://doi.org/10.1017/CBO9781139015516
[7] Chandrasekaran AR, Punnoose JA, Zhou L, Dey P, Dey BK, Halvorsen K (2019) DNA nanotechnology approaches for
microRNA detection and diagnosis. Nucleic Acids Research 47(20):10489–10505. https://doi.org/10.1093/nar/gkz580
[8] Ke Y, Castro C, Choi JH (2018) Structural DNA Nanotechnology : Artificial Nanostructures for Biomedical. 375–403.
[9] Shen Z, Yan H, Wang T, Seeman NC (2004) Paranemic crossover DNA: a generalized Holliday structure with applications
in nanotechnology. Journal of the American Chemical Society 126(6):1666–74. https://doi.org/10.1021/ja038381e
[10] Rothemund PWK (2006) Folding DNA to create nanoscale shapes and patterns. Nature 440(7082):297–302.
https://doi.org/10.1038/nature04586
[11] Woo S, Rothemund PWK (2014) Self-assembly of two-dimensional DNA origami lattices using cation-controlled surface
diffusion. Nature Communications 5:4889. https://doi.org/10.1038/ncomms5889
[12] Woo S, Rothemund PWK (2011) Programmable molecular recognition based on the geometry of DNA nanostructures.
Nature Chemistry 3(8):620–7. https://doi.org/10.1038/nchem.1070
[13] Strong M (2004) Protein Nanomachines. PLOS Biology 2(3). https://doi.org/10.1371/journal.pbio.0020073
[14] Jun H, Shepherd TR, Zhang K, Bricker WP, Li S, Chiu W, Bathe M (2019) Automated sequence design of 3D polyhedral
wireframe DNA origami with honeycomb edges. ACS Nano 13(2):2083–2093. https://doi.org/10.1021/acsnano.8b08671
[15] Jun H, Wang X, Bricker WP, Bathe M (2019) Automated sequence design of 2D wireframe DNA origami with honeycomb
edges. Nature Communications 10(1):1–9. https://doi.org/10.1038/s41467-019-13457-y
[16] Jun H, Zhang F, Shepherd T, Ratanalert S, Qi X, Yan H, Bathe M (2019) Autonomously designed free-form 2D DNA
origami. Science Advances 5(1):1–9. https://doi.org/10.1126/sciadv.aav0655
[17] SantaLucia J, Hicks D (2004) The thermodynamics of DNA structural motifs. Annual Review of Biophysics and
Biomolecular Structure 33:415–440. https://doi.org/10.1146/annurev.biophys.32.110601.141800
[18] Dannenberg F, Dunn KE, Bath J, Turberfield AJ, Ouldridge TE, Kwiatkowska M, Turberfield AJ, Ouldridge TE (2015)
Modelling DNA origami self-assembly at the domain level. The Journal of Chemical Physics 143(16):1–19.
https://doi.org/10.1063/1.4933426
[19] Arbona J-M, Aimé J-P, Elezgaray J (2013) Cooperativity in the annealing of DNA origamis. The Journal of Chemical
Physics 138(1):015105. https://doi.org/10.1063/1.4773405
[20] Crothers DM, Drak J, Kahn JD, Levene SDBT-M in E (1992) DNA bending, flexibility, and helical repeat by cyclization
kinetics. DNA Structures Part B: Chemical and Electrophoretic Analysis of DNA (Academic Press), Vol. 212, pp 3–29.
https://doi.org/10.1016/0076-6879(92)12003-9
[21] Jacobson H, Stockmayer WH (1950) Intramolecular Reaction in Polycondensations. I. The Theory of Linear Systems. The
Journal of Chemical Physics 1600(18). https://doi.org/10.1063/1.1747547
[22] Zhang Y, Mcewen AE, Crothers DM, Levene SD (2006) Statistical-Mechanical Theory of DNA Looping. Biophysical
Journal 90(March):1903–1912. https://doi.org/10.1529/biophysj.105.070490
[23] Yakovchuk P, Protozanova E, Frank-Kamenetskii MD (2006) Base-stacking and base-pairing contributions into thermal
stability of the DNA double helix. Nucleic Acids Research 34(2):564–574. https://doi.org/10.1093/nar/gkj454
[24] Strauss MT, Schueder F, Haas D, Nickels PC, Jungmann R (2018) Quantifying absolute addressability in DNA origami with
molecular resolution. Nature Communications 9(1):1–7.
https://doi.org/10.1038/s41467-018-04031-z
[25] Lee Tin Wah J, David C, Rudiuk S, Baigl D, Estevez-Torres AAA, Wah JLT, David C, Rudiuk S, Baigl D, Estevez-Torres
AAA (2016) Observing and Controlling the Folding Pathway of DNA Origami at the Nanoscale. ACS Nano 10(2):1978–
1987. https://doi.org/10.1021/acsnano.5b05972
[26] Jacobs WM, Reinhardt A, Frenkel D (2015) Rational design of self-assembly pathways for complex multicomponent
structures. Proceedings of the National Academy of Sciences of the United States of America 112(20):6313–8.
https://doi.org/10.1073/pnas.1502210112
[27] Dunn KE, Dannenberg F, Ouldridge TE, Kwiatkowska M, Turberfield AJ, Bath J (2015) Guiding the folding pathway of
DNA origami. Nature. https://doi.org/10.1038/nature14860
[28] Wei X, Nangreave J, Liu Y (2014) Uncovering the self-assembly of DNA nanostructures by thermodynamics and kinetics.
Accounts of Chemical Research 47(6):1861–1870. https://doi.org/10.1021/ar5000665
[29] Owczarzy R, Tataurov A V., Wu Y, Manthey JA, McQuisten KA, Almabrazi HG, Pedersen KF, Lin Y, Garretson J,
McEntaggart NO, Sailor CA, Dawson RB, Peek AS (2008) IDT SciTools: a suite for analysis and design of nucleic acid
oligomers. Nucleic Acids Research 36(Web Server issue):W163-9. https://doi.org/10.1093/nar/gkn198
[30] Kibbe WA (2007) OligoCalc: an online oligonucleotide properties calculator. Nucleic Acids Research 35(Web Server
issue):W43-6. https://doi.org/10.1093/nar/gkm234
[31] Tulpan D, Andronescu M, Leger S (2010) Free energy estimation of short DNA duplex hybridizations. BMC Bioinformatics
11. https://doi.org/10.1186/1471-2105-11-105

Volume 126, Article No. 126001 (2021) https://doi.org/10.6028/jres.126.001
Journal of Research of the National Institute of Standards and Technology
33 https://doi.org/10.6028/jres.126.001
[32] Kim DN, Kilchherr F, Dietz H, Bathe M (2012) Quantitative prediction of 3D solution shape and flexibility of nucleic acid
nanostructures. Nucleic Acids Research 40(7):1–7. https://doi.org/10.1093/nar/gkr1173
[33] Maffeo C, Yoo J, Aksimentiev A (2016) De Novo Reconstruction of DNA Origami Structures through Atomistic Molecular
Dynamics Simulation. Nucleic Acids Research 44:3013–3019. https://doi.org/10.1093/nar/gkw155
[34] Yoo J, Aksimentiev A (2015) Molecular Dynamics of Membrane-Spanning DNA Channels: Conductance Mechanism,
Electro-Osmotic Transport, and Mechanical Gating. The Journal of Physical Chemistry Letters 6:4680–4687.
https://doi.org/10.1021/acs.jpclett.5b01964
[35] Yoo J, Aksimentiev A (2013) In situ structure and dynamics of DNA origami determined through molecular dynamics
simulations. Proceedings of the National Academy of Sciences of the United States of America 110(50):20099–20104.
https://doi.org/10.1073/pnas.1316521110
[36] Jepsen MDE, Sørensen RS, Maffeo C, Aksimentiev A, Kjems J, Birkedal V (2019) Single molecule analysis of structural
fluctuations in DNA nanostructures. Nanoscale 11:18475–18482. https://doi.org/10.1039/C9NR03826D
[37] Doye JPK, Ouldridge TE, Louis AA, Romano F, Šulc P, Matek C, Snodin BEK, Rovigatti L, Schreck JS, Harrison RM,
Smith WPJ (2013) Coarse-graining DNA for simulations of DNA nanotechnology. Physical Chemistry Chemical Physics
15(47):20395–20414. https://doi.org/10.1039/c3cp53545b
[38] Maffeo C, Aksimentiev A (2020) MrDNA: a multi-resolution model for predicting the structure and dynamics of DNA
systems. Nucleic Acids Research. https://doi.org/10.1093/nar/gkaa200
[39] Högberg B, Liedl T, Shih WM (2009) Folding DNA origami from a double-stranded source of scaffold. Journal of the
American Chemical Society 131(26):9154–5. https://doi.org/10.1021/ja902569x
[40] Veneziano R, Shepherd TR, Ratanalert S, Bellou L, Tao C, Bathe M (2018) In vitro synthesis of gene-length single-stranded
DNA. Scientific Reports 8(1):1–7. https://doi.org/10.1038/s41598-018-24677-5
[41] Duffy S, Shackelton LA, Holmes EC (2008) Rates of evolutionary change in viruses: Patterns and determinants. Nature
Reviews Genetics 9(4):267–276. https://doi.org/10.1038/nrg2323
[42] Brown S, Majikes JM, Fennell H, Samano EC, LaBean TH, Martínez A, Girón TM, Fennell H, Samano EC, LaBean TH
(2015) An easy-to-prepare mini-scaffold for DNA origami. Nanoscale 4434(7):16621–16624.
https://doi.org/10.1039/c5nr04921k
[43] Marchi AN, Saaem I, Vogen BN, Brown S, LaBean TH (2014) Towards larger DNA origami. Nano Letters 14(10):5740–
5747. https://doi.org/10.1021/nl502626s
[44] Miao H, De Llano E, Sorger J, Ahmadi Y, Kekic T, Isenberg T, Gröller ME, Barišić I, Viola I (2018) Multiscale
Visualization and Scale-Adaptive Modification of DNA Nanostructures. IEEE Transactions on Visualization and Computer
Graphics 24(1):1014–1024. https://doi.org/10.1109/TVCG.2017.2743981
[45] Benson E, Mohammed A, Gardell J, Masich S, Czeizler E, Orponen P, Högberg B (2015) DNA rendering of polyhedral
meshes at the nanoscale. Nature 523(7561):441–444. https://doi.org/10.1038/nature14586
[46] Veneziano R, Ratanalert S, Zhang K, Zhang F, Yan H, Chiu W, Bathe M (2016) Designer nanoscale DNA assemblies
programmed from the top down. Science 352(6293):1534 LP–1534. https://doi.org/10.1126/science.aaf4388
[47] Ke Y, Bellot G, Voigt N V., Fradkov E, Shih WM (2012) Two design strategies for enhancement of multilayer–DNA-
origami folding: underwinding for specific intercalator rescue and staple-break positioning. Chemical Science 3(8):2587.
https://doi.org/10.1039/c2sc20446k
[48] Baker MAB, Tuckwell AJ, Berengut JF, Bath J, Benn F, Duff AP, Whitten AE, Dunn KE, Hynson RM, Turberfield AJ, Lee
LK (2018) Dimensions and Global Twist of Single-Layer DNA Origami Measured by Small-Angle X-ray Scattering. ACS
Nano 12(6):5791–5799. https://doi.org/10.1021/acsnano.8b01669
[49] Dietz H, Douglas SM, Shih WM (2009) Folding DNA into twisted and curved nanoscale shapes. Science 325(5941):725–
730. https://doi.org/10.1126/science.1174251
About the authors: J. Alexander Liddle is the Scientific Director of the Microsystems and Nanotechnology
Division at NIST. Jacob M. Majikes is a postdoctoral researcher in the Microsystems and Nanotechnology
Division at NIST. The National Institute of Standards and Technology is an agency of the U.S. Department
of Commerce.
