1
January 31
st
2020
Widespread divergent transcription from prokaryotic promoters
Emily Warman
1
, David Forrest
1
, Joseph T. Wade
2,3
, David C. Grainger
1
*
1
Institute for Microbiology and Infection, School of Biosciences, University of Birmingham,
Edgbaston, Birmingham, B15 2TT, UK
2
Wadsworth Centre, New York State Department of Health, Albany, NY, 12208, USA
3
Department of Biomedical Sciences, University at Albany, Albany, NY, 12201, USA
*for correspondence, d.grainger@bham.ac.uk Tel: +44 (0)121 4145437
.CC-BY 4.0 International licenseperpetuity. It is made available under a
preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in
The copyright holder for thisthis version posted February 2, 2020. ; https://doi.org/10.1101/2020.01.31.928960doi: bioRxiv preprint
2
ABSTRACT
Promoters are DNA sequences that stimulate the initiation of transcription. In all prokaryotes,
promoters are believed to drive transcription in a single direction. Here we show that prokaryotic
promoters are frequently bidirectional and drive divergent transcription. Mechanistically, this
occurs because key promoter elements have inherent symmetry and often coincide on opposite
DNA strands. Reciprocal stimulation between divergent transcription start sites also contributes.
Horizontally acquired DNA is enriched for bidirectional promoters suggesting that they represent
an early step in prokaryotic promoter evolution.
Transcription initiation requires DNA sequences called promoters that interact with RNA polymerase
(RNAP)
1
. Promoters consist of ordered core elements with distinct roles
2,3
. For example, most bacterial
promoters contain a -10 element that interacts with the housekeeping RNAP σ
70
subunit. This facilitates
DNA unwinding
4,5
. In eukaryotes and archaea, the TBP binding TATA box has a similar role
6
. It has
long been assumed that promoters are directional, driving transcription in a single orientation
determined by promoter element arrangement
2,7
. This view has recently been challenged in eukaryotes
8–
11
. Nonetheless, the consensus view is that prokaryotic promoters are unidirectional
12
.
Previous studies have mapped transcription start sites (TSSs) in Escherichia coli by detecting
triphosphorylated RNA 5' ends
13
. These TSSs can be assigned to σ
70
binding events identified using
ChIP-seq
13
. We noticed that not all
70
binding sites were associated with detectable RNA 5' ends. This
was particularly evident for horizontally acquired genes silenced by histone-like nucleoid structuring
(H-NS) protein (Figure S1). We reasoned that RNAP might initiate transcription but produce unstable
RNAs. To test this, we fused 33 such
70
targets to lacZ. Any transcripts produced should be stabilised,
and detectable, due to translation. Transcription orientation cannot be directly inferred from
70
ChIP-
seq data. Hence, DNA sequences were cloned in both directions (Figure 1a). Surprisingly, over half of
the fragments were transcriptionally active in both orientations (Figure 1b). We designated the direction
of highest lacZ expression as “forward”. On average, “reverse” transcription neared half the “forward”
activity (Figure 1c). For a subset of divergent transcript pairs, we mapped RNA 5' ends (Figure 1d).
Most reverse TSSs were upstream of the forward TSS and resulted from shared overlapping promoter
elements (Figures 1e and S2). Mutations in shared promoter elements (Figure S2) reduced expression
in both orientations (Figure 1f).
To understand global patterns of divergent transcription we analysed TSSs independently mapped by
RNA 5' polyphosphatase sequencing (PPP-seq), dRNA-seq or cappable-seq
13–15
. In all cases, oppositely
orientated TSSs tended to co-locate (Figure 2a). To increase sensitivity, we merged the datasets (Figure
2a, combined). This identified 5,292 divergent TSSs, defined as being separated by between 25 and 7
.CC-BY 4.0 International licenseperpetuity. It is made available under a
preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in
The copyright holder for thisthis version posted February 2, 2020. ; https://doi.org/10.1101/2020.01.31.928960doi: bioRxiv preprint
3
bp; 19 % of all detected TSSs in E. coli. We refer to the associated promoters as bidirectional. The most
common distance between divergent TSSs was 18 bp; transcription initiates either side of
overlapping -10 elements (Figure 2a, top expansion). We reasoned that promoter element symmetry
must play a major role. To test this, we made a position weight matrix (PWM) describing all E. coli
promoter sequences. If the PWM matched adjacent regions of DNA on opposite strands the symmetry
score increased. Maximum symmetry correlated with divergent transcription (R
2
= 0.85; Figure 2a
bottom expansion). Consistent with this, a DNA sequence logo generated by aligning divergent TSSs,
separated by 18 bp, was symmetrical (Figure 2b). Contrastingly, TSSs with no divergent transcript
generated an asymmetrical motif (Figure 2c). Note that the first, second and sixth positions of promoter
-10 elements (consensus 5'-TATAAT-3') are key for transcription initiation
5
(Figure 2c). At divergent
TSS offset by 18 bp, nucleotides two and six of -10 elements on opposite DNA strands base pair. Hence,
these positions are most strongly conserved (Figure 2b). Example -10 elements arranged in this way are
shown in Figure 2d. Divergent transcription also increased at TSSs separated by 29, 23, 12, 10 or 7 bp
(Figure S3a). These configurations also correspond to symmetrical base pairing between key -10
element nucleotides, and TSSs, on opposite DNA strands (Figure S3b). The distribution of all
bidirectional promoters with respect to genes is shown in Figure 2e.
In E. coli transcription preferentially initiates at an adenine (Figure 2c). For divergent TSSs 18 bp apart,
the +1 nucleotide corresponds to position -18 on the opposite DNA strand. Hence, -18 is often a thymine
(Figure 2b). A thymine at position -18 can increase transcription by improving interaction between
70
residue R451 and the DNA backbone
16
(Figure 3a). We speculated that the +1/-18 overlap could explain
why this configuration is so frequently detected. To test this, we cloned a bidirectional promoter, with
18 bp between TSSs, in both orientations upstream of the oop transcriptional terminator (Figure 3bi).
We also made derivatives where the A•T at each +1/-18 position was replaced with C•G (Figure 3bii-
iii). We measured RNA synthesis terminated by oop using in vitro transcription (Figure 3c). As
expected, altering the TSS reduced production of the associated RNA (compare lane 1 with 5 and 3
with 11); the same mutations also reduced transcription in the opposite direction (compare lane 1 with
9 and 3 with 7). Though σ
70
RA451 was defective at the bidirectional promoters (even lane numbers to
12) it was unimpaired at a control promoter not requiring this contact (lanes 13-14).
To determine the prevalence of bidirectional promoters in bacteria we analysed TSS maps for
proteobacteria
13–15,17–20
, actinobacteria
21,22
, and a firmicute
23
. We also mapped TSSs in an additional
firmicute, Bacillus subtilis, using cappable-seq (summarised in Figure S4 and Table S1). Co-localised
divergent TSSs were abundant in all bacteria analysed (Figure 4a). Proteobacteria and actinobacteria
were most similar; divergent TSSs were most frequently separated by 18 or 19 bp, and shared a near-
identical symmetrical -10 element with E. coli (Figure S5). Firmicutes used the same range of -10
.CC-BY 4.0 International licenseperpetuity. It is made available under a
preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in
The copyright holder for thisthis version posted February 2, 2020. ; https://doi.org/10.1101/2020.01.31.928960doi: bioRxiv preprint
4
element configurations illustrated in Figure S3 for E. coli, albeit with little preference for a single
arrangement (Figure S5). For all bacteria, spacing intervals associated with divergent transcription
scored highest for symmetry (Figure S5)
Archaeal transcription is closely related to that of eukaryotes; promoters have a TATA box and B
recognition element (BRE), located a narrow range of distances from the TSS
24
. We analysed TSS maps
for the archaea Thermococcus kodakarensis and Haloferax volcanii
25,26
. We observed strong signatures
of promoter bidirectionality (Figure 4a). In T. kodakarensis, divergent TSSs were predominantly
separated by 52 bp and located either side of a shared TATA box element (5'-TTATAAA-3') (Figure
4b,c and S6a). Less frequently, TSSs separated by 36 bp were used (Figures 4b and S6a). Here, the B
recognition element (BRE; 5'-CGAAA-3') is positioned so the initial C•G bp can also act as the TSS on
the opposite DNA strand (Figure 4c). Similar observations were made for H. volcanii despite the
unusual TATA box consensus (5'-TTWT-3') of haloarchaea (Figure S6b,c). For both species, an
independent promoter PWM search identified near identical spacing rules (Figure S6).
Our data demonstrate that divergent transcription from promoters is a process conserved in all life
forms. The phenomenon is similarly frequent in diverse prokaryotes (Figure S7) and superficially
resembles the situation in eukaryotes. However, the mechanistic basis is fundamentally different (Figure
4d). In eukaryotes, bidirectionality is generated by an activator protein creating two adjacent regions of
nucleosome depletion
27
. Thus, divergent TSSs use separate core promoter elements that can be
separated by thousands of bp, with no distance optimal. By contrast, divergent transcription in
prokaryotes depends on symmetry of key promoter elements; TSSs on opposite strands are closely
spaced at preferred intervals. Consequently, for prokaryotes, divergent transcription can be predicted
using DNA sequence and recapitulated in vitro with purified components. In eukaryotes, recently
acquired DNA is enriched for bidirectional promoters
27
. This has been attributed to pervasive
transcription factor binding
27
. We initially identified divergent transcription in horizontally acquired E.
coli DNA (Figure 1). Furthermore, detection of bidirectional promoters increased in cells lacking H-
NS (Figure S8a). This results from elevated promoter frequency and symmetry in foreign genes (Figure
S8b). Hence, divergent transcription and promoter evolution are linked in prokaryotes. Strikingly, the
proportion of bidirectional promoters used for mRNA production is higher than the equivalent fraction
of canonical promoters (Figures 2e, S4d). We conclude that divergent transcription plays a key role in
prokaryotic cells.
.CC-BY 4.0 International licenseperpetuity. It is made available under a
preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in
The copyright holder for thisthis version posted February 2, 2020. ; https://doi.org/10.1101/2020.01.31.928960doi: bioRxiv preprint
5
FIGURE LEGENDS
Figure 1: Divergent transcription within horizontally acquired genes. a) -galactosidase activity
derived from cryptic RNAP binding sites. b) Direction of transcription from cloned DNA fragments. c)
Average forward or reverse -galactosidase activity of all DNA fragments. d) Start sites mapped by
primer extension for selected DNA fragments (orientations labelled a or b). Primer extension products
in lanes 1 to 10, sizes in nucleotides (nt). Lanes 11-14 are Maxam-Gilbert sequencing reactions for
calibration. e) Schematic representation of core promoter elements associated with divergent
transcription. f) Effect of mutating shared core promoter elements.
Figure 2: Widespread divergent transcription from bidirectional promoters in Escherichia coli.
a) Heatmaps made using global transcription start site (TSS) data
13–15
or position weight matrix
analysis. TSSs on the top chromosome strand are aligned at the centre of the heatmap (bent arrow,
labelled +1). Heatmap colour indicates abundance of bottom strand TSSs at that position. The expansion
shows the occurrence of bottom strand TSSs in a 50 bp window either side of all top strand promoters.
b) Predominant DNA sequence motif associated with bidirectional or c) directional promoters. The x-
axis break indicates the variable distance between -10 element and TSS at directional promoters. d) a
bidirectional promoter between the E. coli pfs and dgt genes. Promoter -10 elements are bold. TSSs are
in uppercase. e) Relative position of all bidirectional E. coli promoters with respect to genes.
Figure 3: Reciprocal stimulation between divergent transcription start sites. a) Structure of RNAP
bound to DNA (PDB: 6CA0)
28
. Relevant features labelled. b) DNA templates used for in vitro
transcription. For simplicity, only the sequences of promoter -10 elements (labelled) and TSSs (bent
arrows) are shown. Opposing DNA strands represented by black or blue lines. Interaction between
70
R451 and the DNA backbone indicated by dashes. c) Products of in vitro transcription (using templates
in panel b) using either
70
or the R451A derivative. The RNAI transcript is derived from the replication
origin of the plasmid DNA template.
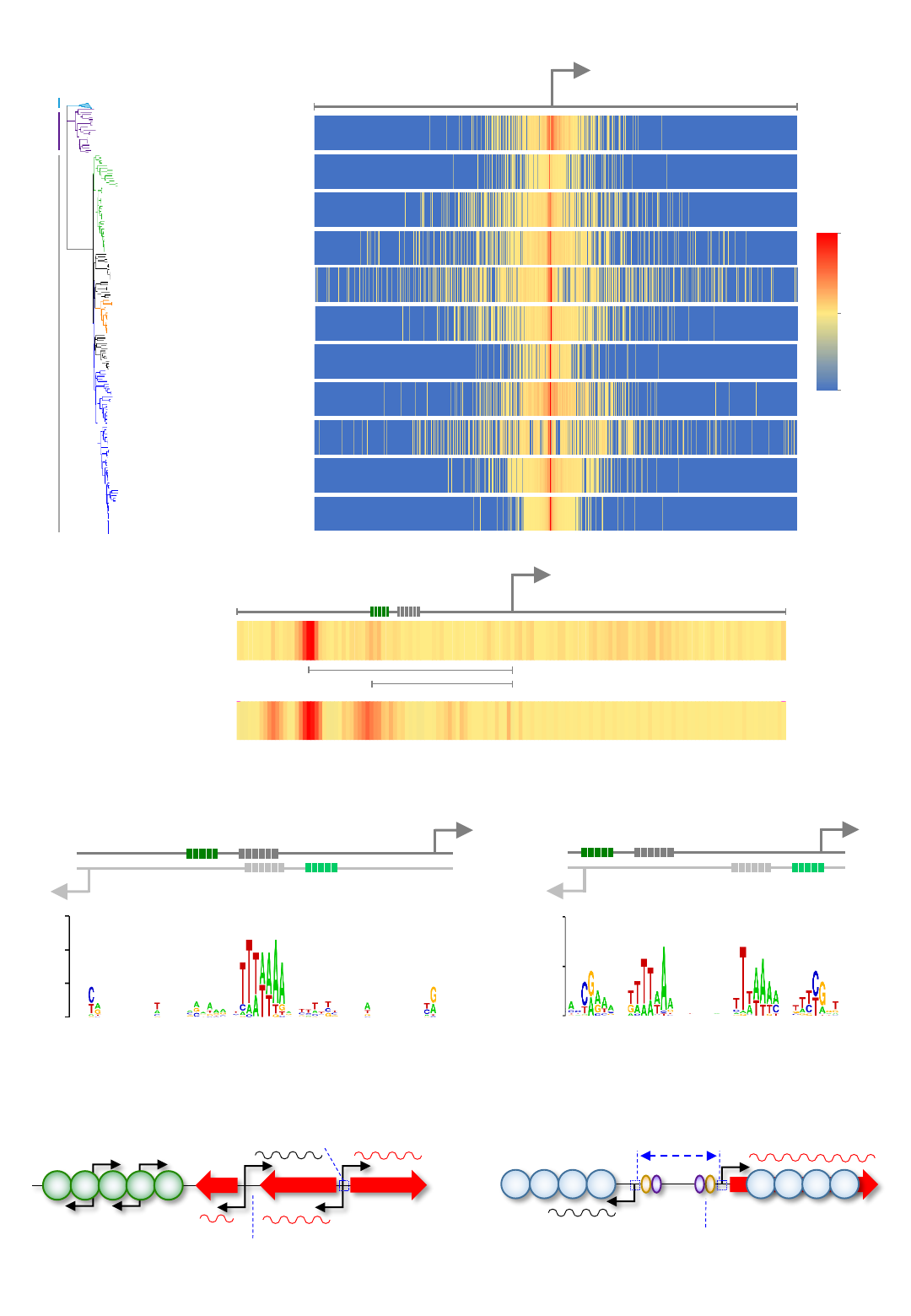
Figure 4: Bidirectional promoters are widespread in prokaryotes. a,b) Heatmaps indicate
abundance and position of TSSs on the bottom DNA strand, relative to the nearest top strand promoter
(bent arrow). Species and phylogenetic relationships are indicated to left of heatmaps. c) DNA sequence
motifs derived from divergent TSSs in T. kodakarensis. d) Bidirectional promoters have a different
basis in prokaryotes and eukaryotes.
REFERENCES
1. Decker, K. B. & Hinton, D. M. Transcription regulation at the core: similarities among
bacterial, archaeal, and eukaryotic RNA polymerases. Annu. Rev. Microbiol. 67, 113–39
(2013).
2. Browning, D. F. & Busby, S. J. W. The regulation of bacterial transcription initiation. Nature
Reviews Microbiology vol. 2 57–65 (2004).
3. Haberle, V. & Stark, A. Eukaryotic core promoters and the functional basis of transcription
initiation. Nature Reviews Molecular Cell Biology vol. 19 621–637 (2018).
4. Bae, B., Feklistov, A., Lass-Napiorkowska, A., Landick, R. & Darst, S. A. Structure of a
bacterial RNA polymerase holoenzyme open promoter complex. Elife 4, (2015).
5. Feklistov, A. & Darst, S. A. Structural basis for promoter -10 element recognition by the
bacterial RNA polymerase σ subunit. Cell 147, 1257–1269 (2011).
6. Kramm, K., Engel, C. & Grohmann, D. Transcription initiation factor TBP: Old friend new
questions. Biochemical Society Transactions vol. 47 411–423 (2019).
.CC-BY 4.0 International licenseperpetuity. It is made available under a
preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in
The copyright holder for thisthis version posted February 2, 2020. ; https://doi.org/10.1101/2020.01.31.928960doi: bioRxiv preprint
6
7. Butler, J. E. F. & Kadonaga, J. T. The RNA polymerase II core promoter: A key component in
the regulation of gene expression. Genes and Development vol. 16 2583–2592 (2002).
8. Core, L. J., Waterfall, J. J. & Lis, J. T. Nascent RNA sequencing reveals widespread pausing
and divergent initiation at human promoters. Science (80-. ). 322, 1845–1848 (2008).
9. Seila, A. C. et al. Divergent transcription from active promoters. Science (80-. ). 322, 1849–
1851 (2008).
10. Preker, P. et al. RNA exosome depletion reveals transcription upstream of active human
promoters. Science (80-. ). 322, 1851–1854 (2008).
11. He, Y., Vogelstein, B., Velculescu, V. E., Papadopoulos, N. & Kinzler, K. W. The antisense
transcriptomes of human cells. Science (80-. ). 322, 1855–1857 (2008).
12. Browning, D. F. & Busby, S. J. W. Local and global regulation of transcription initiation in
bacteria. Nature Reviews Microbiology vol. 14 638–650 (2016).
13. Singh, S. S. et al. Widespread suppression of intragenic transcription initiation by H-NS.
Genes Dev. 28, 214–219 (2014).
14. Thomason, M. K. et al. Global transcriptional start site mapping using differential RNA
sequencing reveals novel antisense RNAs in Escherichia coli. J. Bacteriol. 197, 18–28 (2015).
15. Ettwiller, L., Buswell, J., Yigit, E. & Schildkraut, I. A novel enrichment strategy reveals
unprecedented number of novel transcription start sites at single base resolution in a model
prokaryote and the gut microbiome. BMC Genomics 17, 199 (2016).
16. Singh, S. S., Typas, A., Hengge, R. & Grainger, D. C. Escherichia coli σ 70 senses sequence
and conformation of the promoter spacer region. Nucleic Acids Res. 39, 5109–5118 (2011).
17. Papenfort, K., Förstner, K. U., Cong, J. P., Sharma, C. M. & Bassler, B. L. Differential RNA-
seq of Vibrio cholerae identifies the VqmR small RNA as a regulator of biofilm formation.
Proc. Natl. Acad. Sci. U. S. A. 112, E766–E775 (2015).
18. Gill, E. E. et al. High-throughput detection of RNA processing in bacteria. BMC Genomics 19,
(2018).
19. Kröger, C. et al. The primary transcriptome, small RNAs and regulation of antimicrobial
resistance in Acinetobacter baumannii ATCC 17978. Nucleic Acids Res. 46, 9684–9698
(2018).
20. Sharma, C. M. et al. The primary transcriptome of the major human pathogen Helicobacter
pylori. Nature 464, 250–255 (2010).
21. Cortes, T. et al. Genome-wide Mapping of Transcriptional Start Sites Defines an Extensive
Leaderless Transcriptome in Mycobacterium tuberculosis. Cell Rep. 5, 1121–1131 (2013).
22. Jeong, Y. et al. The dynamic transcriptional and translational landscape of the model antibiotic
producer Streptomyces coelicolor A3(2). Nat. Commun. 7, (2016).
23. Fan, B. et al. dRNA-Seq Reveals Genomewide TSSs and Noncoding RNAs of Plant Beneficial
Rhizobacterium Bacillus amyloliquefaciens FZB42. PLoS One 10, e0142002 (2015).
24. Decker, K. B. & Hinton, D. M. Transcription Regulation at the Core: Similarities Among
Bacterial, Archaeal, and Eukaryotic RNA Polymerases. Annu. Rev. Microbiol. 67, 113–139
(2013).
25. Babski, J. et al. Genome-wide identification of transcriptional start sites in the haloarchaeon
Haloferax volcanii based on differential RNA-Seq (dRNA-Seq). BMC Genomics 17, (2016).
26. Jäger, D., Förstner, K. U., Sharma, C. M., Santangelo, T. J. & Reeve, J. N. Primary
.CC-BY 4.0 International licenseperpetuity. It is made available under a
preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in
The copyright holder for thisthis version posted February 2, 2020. ; https://doi.org/10.1101/2020.01.31.928960doi: bioRxiv preprint
7
transcriptome map of the hyperthermophilic archaeon Thermococcus kodakarensis. BMC
Genomics 15, 684 (2014).
27. Jin, Y., Eser, U., Struhl, K. & Churchman, L. S. The Ground State and Evolution of Promoter
Region Directionality. Cell 170, 889-898.e10 (2017).
28. Narayanan, A. et al. Cryo-EM structure of Escherichia coli 70 RNA polymerase and promoter
DNA complex revealed a role of non-conserved region during the open complex formation. J.
Biol. Chem. 293, 7367–7375 (2018).
.CC-BY 4.0 International licenseperpetuity. It is made available under a
preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in
The copyright holder for thisthis version posted February 2, 2020. ; https://doi.org/10.1101/2020.01.31.928960doi: bioRxiv preprint

0.00
100.00
200.00
300.00
400.00
500.00
600.00
700.00
800.00
a b c
f
Divergent Directional
No activity
Mean activity (%)
0
20
40
60
80
100
forward reverse
cryptic RNA polymerase
binding sites
cryptic RNA polymerase
binding sites
0
20
40
60
80
100
120
0
20
40
60
80
100
120
d e
wzxB
-10
-35
-35
-10
5'
5'3'
3'
yibA2
-35
-10
-10
-35
5'
3' 5'
3'
yigG
-10
-35
-35
-10
5'
3' 5'
3'
yqiI2
-10
-35
-35
-10
5'
3' 5'
3'
ygaQ1
-10
-35
-35
-10
5'
3' 5'
3'
control
wzxB
a b
yigG
a b
yqiI2
a b
wild type
mutant
promoter derivative and orientation
Miller units
0
500
1000
2000
1500
Figure 1
190 nt
180 nt
170 nt
160 nt
150 nt
140 nt
130 nt
120 nt
110 nt
marker
yibA2 wzxB yigG yqiI2 ygaQ1
a b a b a b a b a b
175 nt
174 nt
167 nt
167 nt
153 nt
145 nt
147 nt
160 nt
113 nt
113 nt
Lane 1 142 3 4 5 6 7 8 9 10 11 12 13
177 nt
orientation a
orientation b
0
200
400
600
800
control
yeeL
yigF
yigG
gadE
ybdO1
ybdO2
yehA
fepE
leuO
mcrC1
mcrC2
mcrB1
mcrB2
trkG1
trkG2
trkG3
idnK1
idnk2
yqiI1
yqiI2
ygaQ1
ygaQ2
evgS1
evgS2
evgS3
yibA1
yibA2
elaD
sfmD
yeaI
ycjW
wzxB
yagM
Miller units
cryptic RNA polymerase binding sites
.CC-BY 4.0 International licenseperpetuity. It is made available under a
preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in
The copyright holder for thisthis version posted February 2, 2020. ; https://doi.org/10.1101/2020.01.31.928960doi: bioRxiv preprint

dRNA-seq
cappable-seq
PPP-seq
Position of transcription start site on top DNA strand
minimal
maximal
medial
-11.5 kb +11.5 kb
Figure 2
combined
combined
+1
a
Occurrence of nearest TSS on bottom
DNA strand
b
-10 element
+50 bp-50 bp
d e
+1
5,292 bidirectional promoters
coding divergent
convergentco-oriented
18 bp
atttgaaggcatagtttaccatGcgc
taaacTtccgtatcaaatggtacgcg
pfs
dgt
c
I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
position (bp)
bits
0.0
0.5
1.0
1.5
5' 3'
+1
+1
-10 element
-10 element
divergent TSSs (18 bp spacing)
I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
position (bp)
bits
0.0
0.5
1.0
1.5
5' 3'
+1
-10 element
directional TSSs
RNA (PPP-seq)
prediction derived from
PWM symmetry score
0.248989
0.370507
0.441136
0.864629
1.230075
0.89517
0.611122
0.508459
0.236086
0.310393
0.754229
0.9669
0.956196
0.812588
0.411732
0.459451
0.498259
0.142801
0.337937
0.731508
0.713604
0.821426
1.000727
0.57071
0.465336
0.682358
0.827218
0.977018
1.002344
0.825564
0.706353
1.724325
1.984006
1.705626
1.752731
1.048796
0.033982
0.933821
0.800316
0.284779
1.470984
0.147685
0.244769
1.152378
0.211985
0.284147
0.752716
-0.12175
0.125184
0.84214
-0.67402
0.549279
0.423324
0.142136
0.223749
0.401411
0.224748
0.204898
0.320032
0.173048
0.181098
0.285087
0.196466
0.183618
0.348963
0.243064
0.205549
0.358263
0.219576
0.198524
0.315745
0.203804
0.195289
0.344746
0.20631
0.223278
0.356107
0.211141
0.224456
0.308702
0.191761
0.171716
0.296127
0.167652
0.191963
0.315748
0.189042
0.187558
0.293464
0.176466
0.152181
0.265124
0.122686
0.136394
0.252222
0.155648
0.179004
0.277703
0.157529
0.140231
.CC-BY 4.0 International licenseperpetuity. It is made available under a
preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in
The copyright holder for thisthis version posted February 2, 2020. ; https://doi.org/10.1101/2020.01.31.928960doi: bioRxiv preprint

s
70
b
b
'
a
w
Figure 3
a b
R451
-18
-17
c
s
70
RA451
WT
RA451
WT
RA451
WT
RA451
WT
RA451
WT
RA451
WT
RA451
WT
Lane
1
2 3 4 5 6 7 8 9 10 11 12 13
14
controli(f) i(r) ii(f) ii(r) iii(f) iii(r)
RNAI
control
or
invert DNA orientation
i(f)
(r)
ii(f)
(r)
iii(f)
(r)
invert DNA orientation
invert DNA orientation
TATTAT
A
T
TAATAT
A
T
R451
s
70
TATAAT
A
T
TATTAT
A
T
s
70
R451
TATTAT
C
G
TAATAT
A
T
R451
s
70
TATAAT
A
T
TATTAT
C
G
s
70
R451
TATTAT
A
T
TAATAT
C
G
R451
s
70
TATAAT
C
G
TATTAT
A
T
s
70
R451
-10 elements
.CC-BY 4.0 International licenseperpetuity. It is made available under a
preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in
The copyright holder for thisthis version posted February 2, 2020. ; https://doi.org/10.1101/2020.01.31.928960doi: bioRxiv preprint
I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
a
d
minimal
maximal
mediial
Occurrence of
nearest TSS
on bottom
DNA strand
Escherichia coli
11
Vibrio cholerae
10
Pseudomonas aeruginosa
9
Helicobacter pylori
7
Acinetobacter baumannii
8
Mycobacterium tuberculosis
6
Streptomyces coelicolor
5
Bacillus amyloliquefaciens
3
Position of TSS on top DNA strand
+1
-11.5 kb
+11.5 kb
Haloferax volcanii
1
Thermococcus kodakarensis
2
eukaryotes
nucleosome depleted/modified region
prokaryotes
horizontally
acquired region
overlapping sequences/fixed spacing
independent sequences/variable spacing
mRNA
ncRNA
requirement for
transcription factors
Bacillus subtilis
4
Figure 4
11
10
8
9
7
6
5
4
3
1
2
proteobacteria
actinobacteria
firmicutes
bacteria archaea eukarya
b
transcription factor independent
+70 bp
c
+1
TATA boxBRE
+1
TATA box BRE
T. kodakarensis divergent TSSs (52 bp spacing)
position (bp)
5' 3'
bits
0.0
0.5
1.0
1.5
+1
TATA boxBRE
+1
TATA box BRE
T. Kodakarensis divergent TSSs (36 bp spacing)
I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
bits
0.0
1.0
2.0
position (bp)
5'
3'
+1
TATA box
-70 bp
T. kodakarensis
BRE
52 bp
36 bp
H-NS histones
prediction derived from
PWM symmetry score
-0.67376
-1.33736
-1.17406
-0.4816
-0.59024
-0.35034
0.604433
1.222764
2.330164
3.100296
2.343915
1.426643
0.536674
-0.66381
-0.51444
1.055566
2.891743
5.095279
6.589232
6.008972
4.449007
2.584289
0.177109
-1.29526
-1.49504
-1.14171
-0.11193
0.821661
0.511375
0.730977
1.783132
2.268834
2.931526
3.75593
3.073914
2.469459
2.313222
1.058407
0.778709
1.496113
0.800035
0.351962
0.505558
-0.37971
-0.91232
-0.46274
-1.21938
-1.08384
-0.17614
-0.21317
-0.43306
0.386746
0.253954
0.637744
1.10005
0.464467
0.03554
0.992484
0.585699
0.089571
-0.08665
-0.21132
-0.14576
0.197481
-0.31376
-0.09541
0.47873
-0.38123
-0.30457
1.400365
-0.90824
0.013543
0.607924
-0.21625
-0.49171
0.184712
-0.28883
-0.31961
0.095451
-0.36943
-0.3977
0.118542
-0.25899
-0.37207
0.045984
-0.16649
-0.35287
0.064181
-0.18588
-0.26006
0.107061
-0.11442
-0.08557
0.174323
-0.06636
-0.13997
0.099248
-0.18901
-0.35348
-0.0648
-0.23783
-0.24216
0.131807
-0.08823
-0.13318
0.23034
-0.1149
-0.18633
0.08562
-0.20065
-0.15747
0.153394
-0.11419
-0.21143
0.084445
-0.16381
-0.23218
0.001339
-0.24398
-0.28683
0.041066
-0.2086
-0.28081
0.035119
-0.26553
-0.24599
0.033586
-0.08224
-0.11363
0.161791
-0.11853
-0.1951
-0.05035
-0.35915
-0.42289
-0.11571
-0.3459
-0.25599
0.142368
-0.06492
.CC-BY 4.0 International licenseperpetuity. It is made available under a
preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in
The copyright holder for thisthis version posted February 2, 2020. ; https://doi.org/10.1101/2020.01.31.928960doi: bioRxiv preprint
